Circulating SARS-CoV-2 Vaccine Antigen Detected in the Plasma of mRNA-1273 Vaccine Recipients
Alana F. Ogata†1,2,3, Chi-An Cheng†1,2,3, Michaël Desjardins†3,4,5, Yasmeen Senussi1, Amy C. Sherman3,4, Megan Powell4, Lewis Novack4, Salena Von4, Xiaofang Li6, Lindsey R. Baden*3,4,6, David R. Walt*1,2,3
- Department of Pathology, Brigham and Women’s Hospital, Boston, MA, USA
- Wyss Institute for Biologically Inspired Engineering, Harvard University, Boston, MA, USA
- Harvard Medical School, Boston, MA, USA
- Division of Infectious Diseases, Brigham and Women’s Hospital, Boston, MA, USA
- Division of Infectious Diseases, Centre Hospitalier de l’Université de Montréal, Montreal, QC, Canada
- Center for Clinical Investigation, Brigham and Women’s Hospital, Boston, MA, USA
*These authors contributed equally. †These authors contributed equally
- Corresponding author:
David R. Walt
email: dwalt@bwh.harvard.edu
ABSTRACT
SARS-CoV-2 proteins were measured in longitudinal plasma samples collected from 13 participants who received two doses of mRNA-1273 vaccine. 11 of 13 participants showed detectable levels of SARS-CoV-2 protein as early as day one after first vaccine injection. Clearance of detectable SARS-CoV-2 protein correlated with production of IgG and IgA.
KEYWORDS: COVID-19, mRNA vaccine, SARS-CoV-2 antigens, immune responses, spike
INTRODUCTION
Messenger RNA (mRNA) vaccines for coronavirus disease 2019 (COVID-19) have been widely deployed in the US and are highly effective at inducing protection against infection and severe disease1. In December 2020, the Food and Drug Administration gave emergency use authorization (EUA) to the mRNA-1273 vaccine (ModernaTX, Inc), as a two 100 μg dose regimen 28 days apart. mRNA-1273 encodes the severe acute respiratory syndrome coronavirus (SARS-CoV-2) spike antigen with a transmembrane anchor and S1-S2 cleavage site2. Data have demonstrated the elicitation of binding and neutralization antibodies against the spike protein by the mRNA-1273 vaccine in humans, thereby inferring that spike protein was produced and induced an immune response2–5. However, critical data demonstrating the direct production of spike protein via translation from the mRNA-1273 vaccine in these studies are missing, precluding a full understanding of the vaccine mechanism.
Here, we provide evidence that circulating SARS-CoV-2 proteins are present in the plasma of participants vaccinated with the mRNA-1273 vaccine. We report antigen and serological data of the mRNA-1273 vaccine in 13 healthcare workers at the Brigham and Women’s Hospital. Ultrasensitive single-molecule array (Simoa) assays were used for the detection of SARS-CoV-2 antigens spike (S1-S2 unit), S1, and nucleocapsid and antibodies IgG, IgA, and IgM against SARS-CoV-2 spike, S1, receptor binding domain (RBD), and nucleocapsid, as previously described6,7. The ultralow detection limits of the Simoa assays enable detection of antigen and antibody production in the early stages post vaccination and quantification of changes in levels over the course of both vaccine injections.
MATERIALS AND METHODS
A prospective pilot study of 13 healthcare workers, 18 years and older, with no known history of SARS-CoV-2 infection was conducted at the Brigham and Women’s Hospital from December 2020 to March 2021.Full description of study design reported in the Supplementary Materials. SARS-CoV-2 antigens and antibodies were measured using single- molecule array assays, as described in the Supplemental Materials (Methods, Figure S1, Table S2- S4).
RESULTS
Plasma was collected from 13 participants at 10-13 timepoints between 1 and 29 days after the first injection and 1-28 days after the second injection. There was an equal balance of sex, median age was 24 years old, and participants from varied racial and ethnic groups were included (Supplemental Table S1). None of the participants reported prior COVID-19 infection.
Temporal profiling of SARS-CoV-2 antigens and antibodies were acquired using Simoa assays as previously described6,7, providing data on 15 markers to monitor antigen production and immune responses on each participant (Supplemental Figure S2 to S14). The Simoa antigen assays for spike, S1, and nucleocapsid were previously validated in plasma from pre-pandemic healthy subjects, pre-pandemic patients with respiratory infections, COVID-19 negative patients, and COVID-19 positive patients.7 Authors detected S1 and N in 64% of COVID-19 positive patients and S1 levels were significantly associated with disease severity. Here, antigens S1 and spike were measured to probe mRNA translation, while nucleocapsid antigen served as a negative control (Figure 1A, B, C, Supplemental Table S5). After the first 100 μg dose, the mRNA-1273 vaccine produced detectable levels of S1 antigen
in plasma in 11 participants and spike antigen was detected in three of 13 participants. Nucleocapsid antigen was undetectable or at background levels in all participants after both injections, as expected.
S1 antigen was detected as early as day one post vaccination and peak levels were detected on average five days after the first injection (Figure 1A). The mean S1 peak levels was 68 pg/mL ±21 pg/mL. S1 in all participants declined and became undetectable by day 14. No antigen was detected at day zero for 12 of 13 participants, as expected. However, one individual presented detectable S1 on day zero, possibly due to assay cross reactivity with other human coronaviruses or asymptomatic infection at the time of vaccination. Spike protein was detectable in three of 13 participants an average of 15 days after the first injection. The mean spike peak level was 62 pg/mL ± 13 pg/mL. After the second vaccine dose, no S1 or spike was detectable, and both antigens remained undetectable through day 56. For one individual (Participant #8), spike was detected at day 29, one day after the second injection and was undetectable two days later.
Plasma antibodies IgG, IgA, and IgM were measured against spike, S1, RBD, and nucleocapsid. Antibody levels measured on day zero represent the baseline level in each participant, all who had no previous report of COVID-19 infection. In all 13 participants, as expected, IgG levels against spike and S1, and RBD increased after the first injection while IgG against nucleocapsid showed no change over time (Figures 1D, E, F; individual participant data are shown in Supplemental Figures S2-S14). IgA is involved in early neutralization activity and is therefore crucial to target potentially short-lived IgA responses8. Our Simoa assays detected increased IgA against spike, S1, and RBD after the first injection (Supplemental Figure S2-S15). Nine participants (such as participants #3, #4, #5, and #7,
Figure 1G) presented measurable IgG levels against S1 and spike by day 14 after the first vaccine injection. Four participants (participants #1, #6, #10, #13, Supplemental Figure S2, S7, S11, and S14), showed a delayed IgG-S1 and IgG-spike response, which did not increase until day 28 after the first injection. Nonetheless, all participants showed additional boost in IgG-S1 after the second injection.
DISCUSSION
In this study, 11 participants exhibit S1 antigen in plasma after the first injection, while nucleocapsid concentrations are insignificant in all participants, confirming that the detected S1 originates from vaccination and not natural infection. The presence of S1 is likely due to the nature of the encoded mRNA-1273 spike protein, which contains a cleavable S1- S2 site and enables release of S1 from the spike trimer2. We hypothesize that release of S1 protein could result from cleavage via mammalian cell proteases or circulating proteases. We observe an increase in S1 over an initial period of one to five days, suggesting that mRNA translation begins immediately after vaccine inoculation. Interestingly, spike protein appears in three of thirteen participants on average eight days after S1 is produced. The Simoa antigen assays for the full spike protein are designed to require antibody binding to both the S1 and S2 subunits for detection, resulting in a cleaved spike protein to be undetectable. Additionally, spike protein concentrations in plasma of vaccinated participants may be below our assay limit of detection. We hypothesize that the cellular immune responses triggered by T-cell activation, which would occur days after the vaccination, lead to direct killing of cells presenting spike protein and an additional release of spike into the blood stream9. The mechanisms underlying release of free S1 and the subsequent detection of the intact spike protein remain unclear and require further studies.
We additionally present corresponding antibody data that are consistent with serological studies done in large trials2–5,10–12 to support our antigen results. Here, the Simoa serological assays are sensitive enough to identify the production of antibodies at early stages post vaccination and to profile antibody dynamics for each participant at high resolution. IgG and IgA against S1, spike, and RBD increase after S1 production in all participants. There is no significant increase in IgG and IgA against nucleocapsid, confirming that the immune response was specific to the vaccine, which does not contain mRNA for nucleocapsid. For all participants, the increase in IgG against S1 and spike directly corresponds to the decline in S1 or spike protein by the second injection. The inverse correlation between antibody and antigen levels observed is consistent with previous studies investigating SARS-CoV-2 natural infection, in which patients with severe COVID-19 with high plasma antigen levels exhibited antigen clearance upon production of antibodies7. Our Simoa antigen assays cannot detect antigen-antibody immune complexes. Although S1 protein is present in most participants, serological data shows that some participants exhibit an initial enhancement in antibody levels by day 14 compared to those who do not show an enhancement until day 28. These differences in antibody dynamics could be explained by early antibody production due to previous asymptomatic infection, which has recently been demonstrated in seropositive participants10. Here, we observe two participants (Participants #3, #4) with high IgG-spike baseline levels compared to all other participants who produce increased IgG-spike levels by day 14.
Limitations of the current study include the small sample size and potential biases that result from enrolling healthy, young adults, which may not be representative of the general population. Future studies should also examine the dynamics of antigen production with neutralization antibodies. Nonetheless, evidence of systemic detection of spike and S1 protein production from the mRNA-1273 vaccine is significant and has not yet been described in any vaccine study, likely due to limitations in assay sensitivity and timing assessment. The clinical relevance of this finding is unknown and should be further explored. These data show that S1 antigen production after the initial vaccination can be detected by day one and is present beyond the site of injection and the associated regional lymph nodes. Induction of IgG and IgA immune responses can be detected as early as day five post vaccination and are associated with clearance of spike and S1 antigen in the systemic circulation.
NOTES
-
AUTHOR CONTRIBUTIONS
Conceptualization: DRW, AFO, LRB, MD
Investigation: AFO, CC, MD
Sample / Data Acquisition: AFO, CC, MD, YS, ACS, MP, LN, XL, SV Funding acquisition: DRW, LRB
Supervision: DRW, LRB
Writing – original draft: AFO, CC, DRW
Writing – review & editing: MD, ACS, LRB
ACKNOWLEDGMENTS
We thank Jonathan Krauss and John Alexander Kupelian for their technical support and Julia Klopfer for her assistance in sample collections and study management.
FUNDING
Funding for this work came from a generous donation from Barbara and Amos Hostetter and the Chleck Foundation. This work was supported in part by the Bill and Melinda Gates Foundation (no. INV-017380). We acknowledge support from the following research grants for vaccine development: NIH/NIAID, Welcome Trust, and IAVI.
CONFLICTS OF INTEREST
David R. Walt is a Board Director and holds stock equity in Quanterix Corporation, a company that develops an ultra-sensitive digital immunoassay platform. He is an inventory of the Simoa technology, founder of the company and serves on its Board of Directors. Dr. Walt’s interests were reviewed and are managed by Brigham and Women’s Hospital and Partners HealthCare in accordance with their conflict-of-interest policies. The anti-SARS- CoV-2 Simoa assays in this publication have been licensed by Brigham and Women’s Hospital to Quanterix Corporation. A.F.O. reports funding from NIH Post Doctoral Training Grant and royalty payments from Brigham and Women’s Hospital for intellectual property ofthe SARS-CoV-2 antibody assays that was licensed to Quanterix Inc, outside the submitted work. L.R.B. reports research grants from NIH/NIAID, Welcome Trust, and IAVI; and participates in SMCs for NIAID for a variety of programs including vaccine development, outside the submitted work. M.D. reports grants from Centre Hospitalier de l’Université de Montréal (CHUM) and CHUM Foundation, outside the submitted work.
REFERENCES
-
Baden LR, El Sahly HM, Essink B, et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N Engl J Med, 2021; 384: 403-416.
-
Jackson LA, Anderson EJ, Rouphael NG, et al. An mRNA Vaccine against SARS- CoV-2 — Preliminary Report. N Engl J Med, 2020; 383: 1920-1931.
-
Anderson EJ, Rouphael NG, Widge AT, et al. Safety and Immunogenicity of SARS- CoV-2 mRNA-1273 Vaccine in Older Adults. N Engl J Med, 2020; 383: 2427-2438.
-
Widge AT, Rouphael NG, Jackson LA, et al. Durability of Responses after SARS- CoV-2 MRNA-1273 Vaccination. N Engl J Med, 2021; 384: 80-82.
-
Saadat S, Tehrani ZR, Logue J, er al. Binding and Neutralization Antibody Titers After a Single Vaccine Dose in Health Care Workers Previously Infected With SARS-CoV- 2. J Am Med Assoc, 2021; DOI: 10.1001/jama.2021.3341.
-
Norman M, Gilboa T, Ogata AF, et al. Ultrasensitive High-Resolution Profiling of Early Seroconversion in Patients with COVID-19. Nat Biomed Eng, 2020; 4: 1180- 1187.
-
Ogata AF, Maley AM, Wu C, et al. Ultra-Sensitive Serial Profiling of SARS-CoV-2 Antigens and Antibodies in Plasma to Understand Disease Progression in COVID-19 Patients with Severe Disease. Clin Chem, 2020; 4: 1180-1187.
-
Sterlin D, Mathian A, Miyara M, et al. IgA Dominates the Early Neutralizing Antibody Response to SARS-CoV-2. Sci Transl Med, 2021; 13: 2223.
-
Sewell HF, Agius RM, Kendrick D, Stewart M. Covid-19 Vaccines: Delivering Protective Immunity. BMJ, 2020; 371: m4838.
-
Krammer F, Srivastava K, Alshammary H, et al. Antibody Responses in Seropositive Persons after a Single Dose of SARS-CoV-2 mRNA Vaccine. N Engl J Med, 2021; 1- 2.
-
Wu K, Werner AP, Moliva JI, et al. mRNA-1273 Vaccine Induces Neutralizing Antibodies against Spike Mutants from Global SARS-CoV-2 Variants. BioRxiv [Preprint]. January 25, 2021. Available from: https://doi.org/10.1101/2021.01.25.427948.
-
Corbett KS, Edwards DK, Leist SR, et al. SARS-CoV-2 mRNA Vaccine Design Enabled by Prototype Pathogen Preparedness. Nature, 2020; 86: 567-571.
FIGURE LEGENDS
Figure 1.
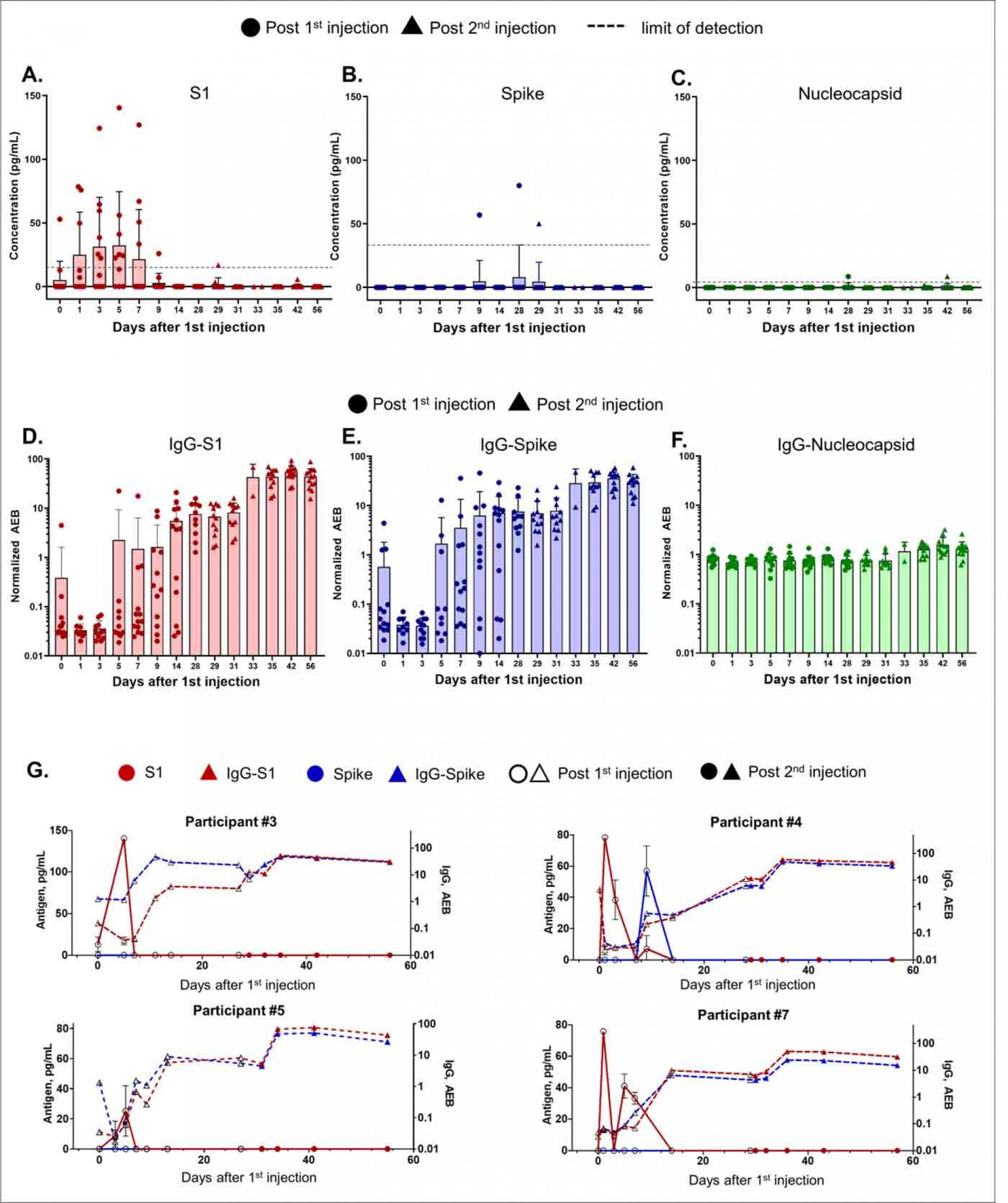
Time Course of SARS-CoV-2 Antigen and Antibody Levels after mRNA-1273 Vaccination. Data from 13 participants. All participants received the mRNA-1273 injection on days 1 and 28, where circle and triangle data points represent antigen levels post the first and second injections, respectively.
(A) S1, (C) spike, and (C) nucleocapsid levels shown were measured using Simoa assays. The dotted line shows the sample limit of detection, calculated as described in the methods. IgG levels against (D) S1, (E) spike, and (F) nucleocapsid shown were measured using Simoa assays and reported in units of normalized Average Enzymes per Bead (AEB). Sample limit of detection in unit of AEB was 0.006 for IgG-S1, 0.0044 for IgG-spike, and 0.0129 for IgG-Nucleocapsid. (A-F) Bars represent the average concentration from participants on that day, and error bars represent the standard deviation, and individual data points are overlayed. Each data point is the average of duplicate measurements. (G) S1, spike, IgG-S1, and IgG-spike data for individual participants #3, #4, #5, and #7. Hollow and solid data points represent levels post the first and second injections, respectively. Each data point represents the average of duplicate measurements, and error bars are the standard deviation.
Accepted Manuscript: Downloaded from Oxford Academic by CovfefeOps on 22 May 2021 Mirrored at this Covfefe Bakery cafe here. Copyright ©2021 The Authors, All Rights Reserved. Published by Oxford University Press for the Infectious Diseases Society of America. All rights reserved. For permissions, e-mail: journals.permissions@oup.com




Comments
Radioactivity